See our Publications tab for a more comprehensive list of papers and preprints. Our research is supported by funding from the NIH (NIBIB and NIA) and NSF.
Brain networks are typically modeled as interactions between neural elements. We have developed a framework for generating networks based on the interactivity of edges. This enables us to investigate circuit-level interactions with exquisite temporal resolution.
Structural connectivity represents the brain’s physical wires — axonal projections or white-matter tracts — whose network organization shapes the correlation structure of neural activity, i.e. functional connectivity. Our work in this area seeks to understand this relationship in greater detail and takes advantage of both in silico simulations and empirical observations.
What routing policy does the brain use to transmit signals and information from one brain area to another? Can these policies be passive and decentralized or do they require top-down, centralized control? We simulate different routing policies, which allows us to explore their advantages and disadvantages, identify tradeoffs, and to develop theories of large-scale inter-areal communication.
Network controllability is a theoretical framework for investigating how time-varying input signals influence the trajectory of a networked dynamical system as it moves through a high-dimensional state space. We apply this framework to biological neural networks to gain insight into the topological properties that support transitions between distinct network states and the energetic cost of driving the network along a desired trajectory towards a target state.
Brain networks grow and evolve over time. This developmental process is the result of a multitude of factors that combine to generate a mature network with functionally adaptive topological properties. We aim to invert this process and uncover the wiring rules and organizational principles that guide brain network development. To this end, we build generative models that realize simple mechanisms of brain network growth and compare the synthetic networks generated by models with empirical observations.
The correlation structure of brain activity fluctuates over short timescales. We use multi-layer network models and time series analysis to investigate the origins of these fluctuations and to assess their relationship with cognitive and psychological processes.
Brain networks are spatially-embedded and the formation, maintenance, and usage of their connections requires material and energy — i.e. costs. In general, longer connections are most costly than short. As a result, brain networks favor forming short-range connections. However, long-distance connections enhance network diversity and promote efficient information transfer. Therefore, brain networks must tradeoff between the formation of costly features that enhance functionality and a brain-wide drive to reduce wiring cost.
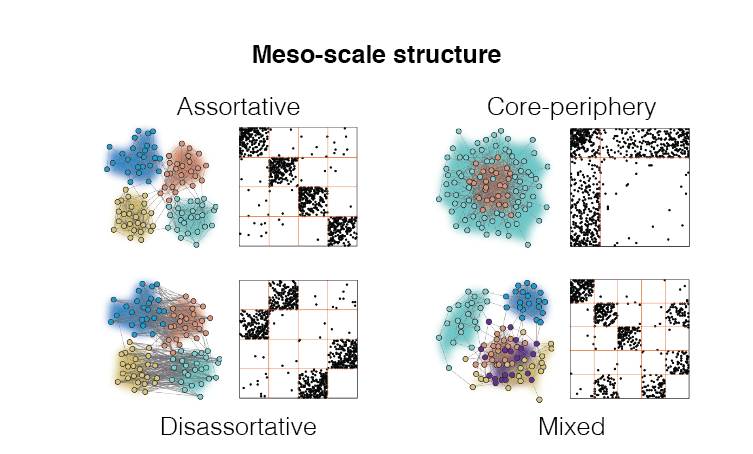
Most real-world networks exhibit meso-scale structure meaning that their nodes and edges can be clustered into sub-networks called “communities”. Communities interact to form different motifs, e.g. assortatively or as cores and peripheries, each suited to realize a different network function. We study meso-scale structure in biological neural networks to gain insight into their relationship with brain function. We also develop new methods for detected and characterizing meso-scale structure.